Characterization of WY 14,643 and its Complex with Aldose Reductase.
Sawaya, M.R., Verma, M., Balendiran, V., Rath, N.P., Cascio, D., Balendiran, G.K.(2016) Sci Rep 6: 34394-34394
- PubMed: 27721416
- DOI: https://doi.org/10.1038/srep34394
- Primary Citation of Related Structures:
5HA7 - PubMed Abstract:
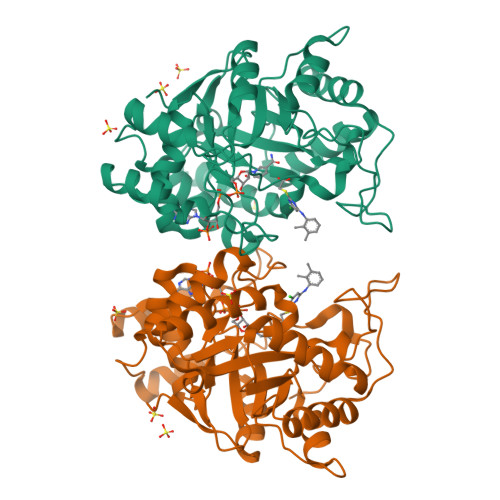
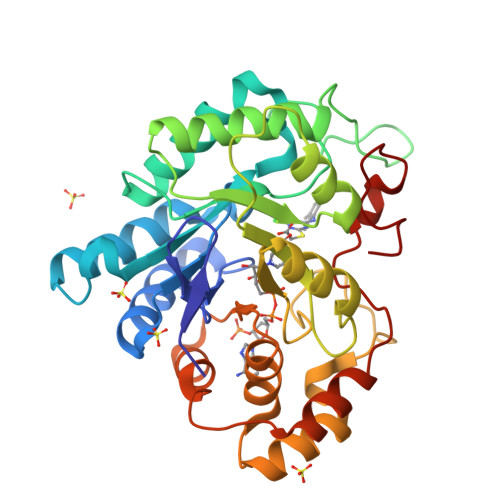
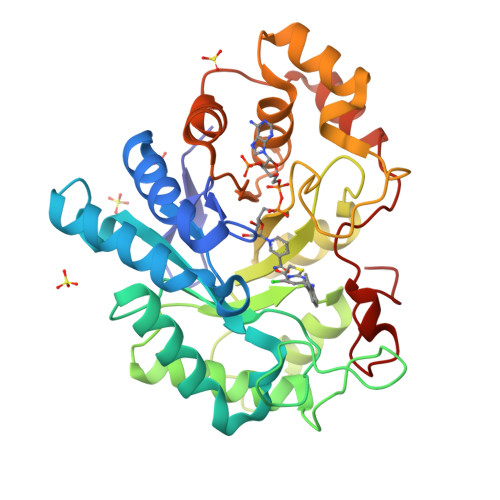
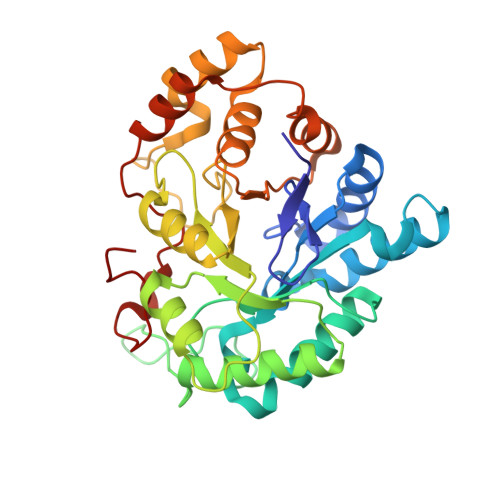
The peroxisome proliferator, WY 14,643 exhibits a pure non-competitive inhibition pattern in the aldehyde reduction and in alcohol oxidation activities of human Aldose reductase (hAR). Fluorescence emission measurements of the equilibrium dissociation constants, K d , of oxidized (hAR•NADP + ) and reduced (hAR•NADPH) holoenzyme complexes display a 2-fold difference between them. K d values for the dissociation of WY 14,643 from the oxidized (hAR•NADP + •WY 14,643) and reduced (hAR•NADPH•WY 14,643) ternary complexes are comparable to each other. The ternary complex structure of hAR•NADP + •WY 14,643 reveals the first structural evidence of a fibrate class drug binding to hAR. These observations demonstrate how fibrate molecules such as WY 14,643, besides being valued as agonists for PPAR, also inhibit hAR.
Organizational Affiliation:
UCLA-DOE, 611 Charles E. Young Drive East, 220 Boyer Hall, Los Angeles, CA 90095, USA.